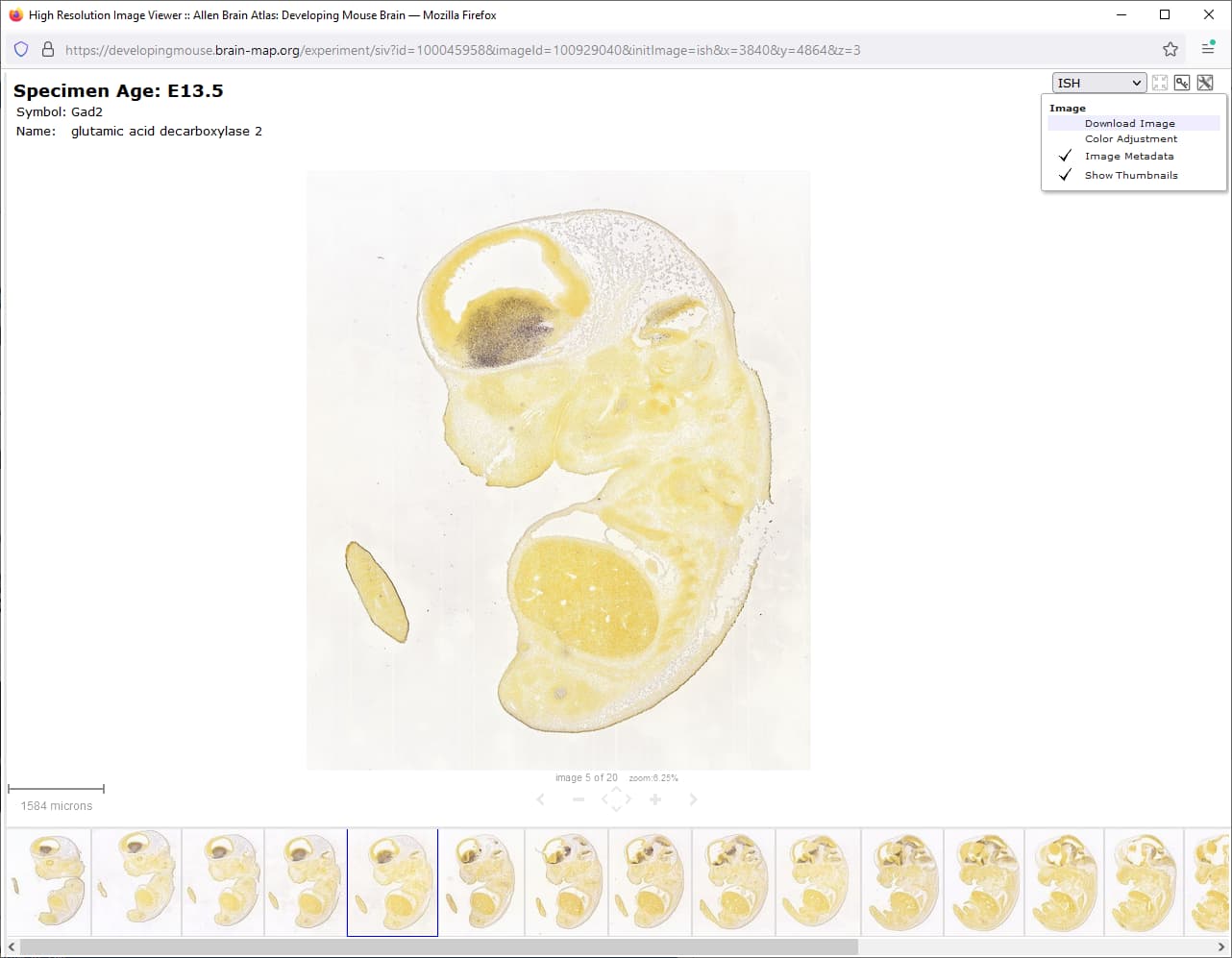
A High-Resolution Spatiotemporal Atlas of Gene Expression of the Developing Mouse Brain - ScienceDirect
The Allen Institute for Brain Science is a 501 (c) (3) nonprofit medical research organization based in Seattle, Washington

ABBA with allen ccfv3 and the Unified Mouse atlas from the Kim lab - Usage & Issues - Image.sc Forum

AllenDigger – spatial expression data visualization, spatial heterogeneity delineation, and single-cell registration based on the Allen Brain Atlas | RNA-Seq Blog

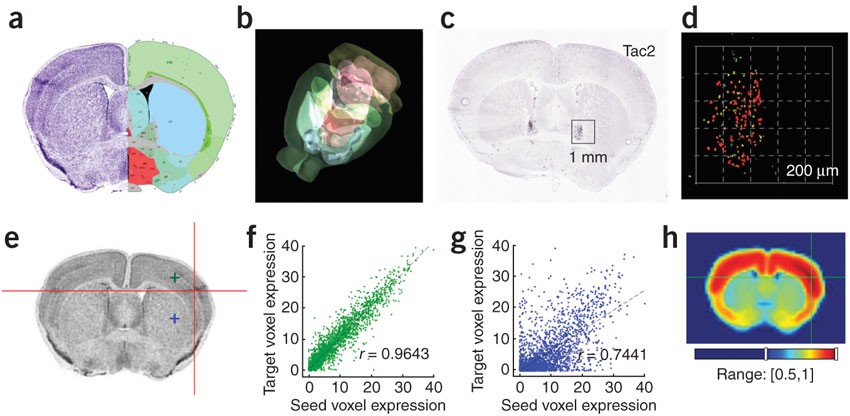
A High-Resolution Spatiotemporal Atlas of Gene Expression of the Developing Mouse Brain - ScienceDirect

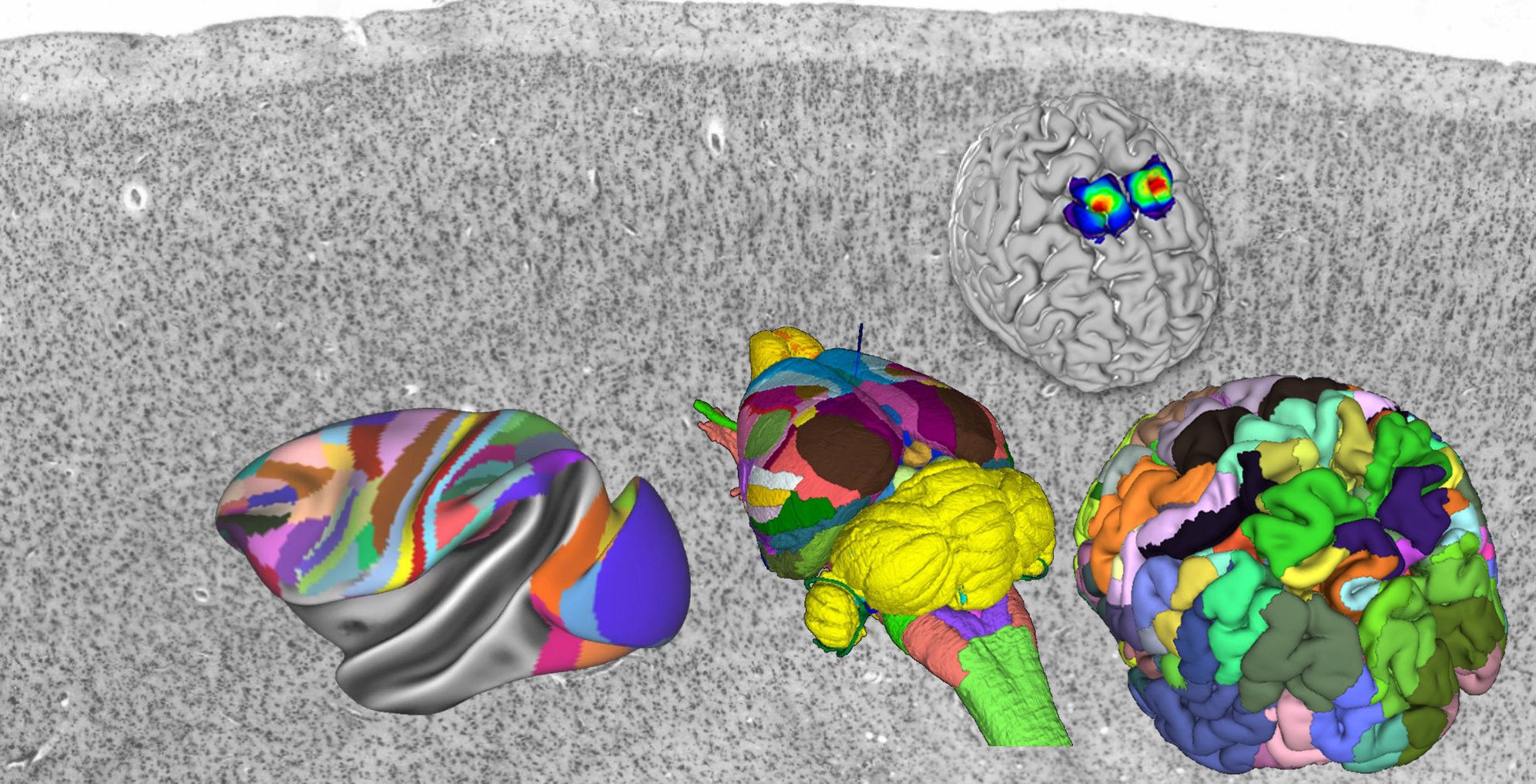
Constructing and optimizing 3D atlases from 2D data with application to the developing mouse brain | eLife
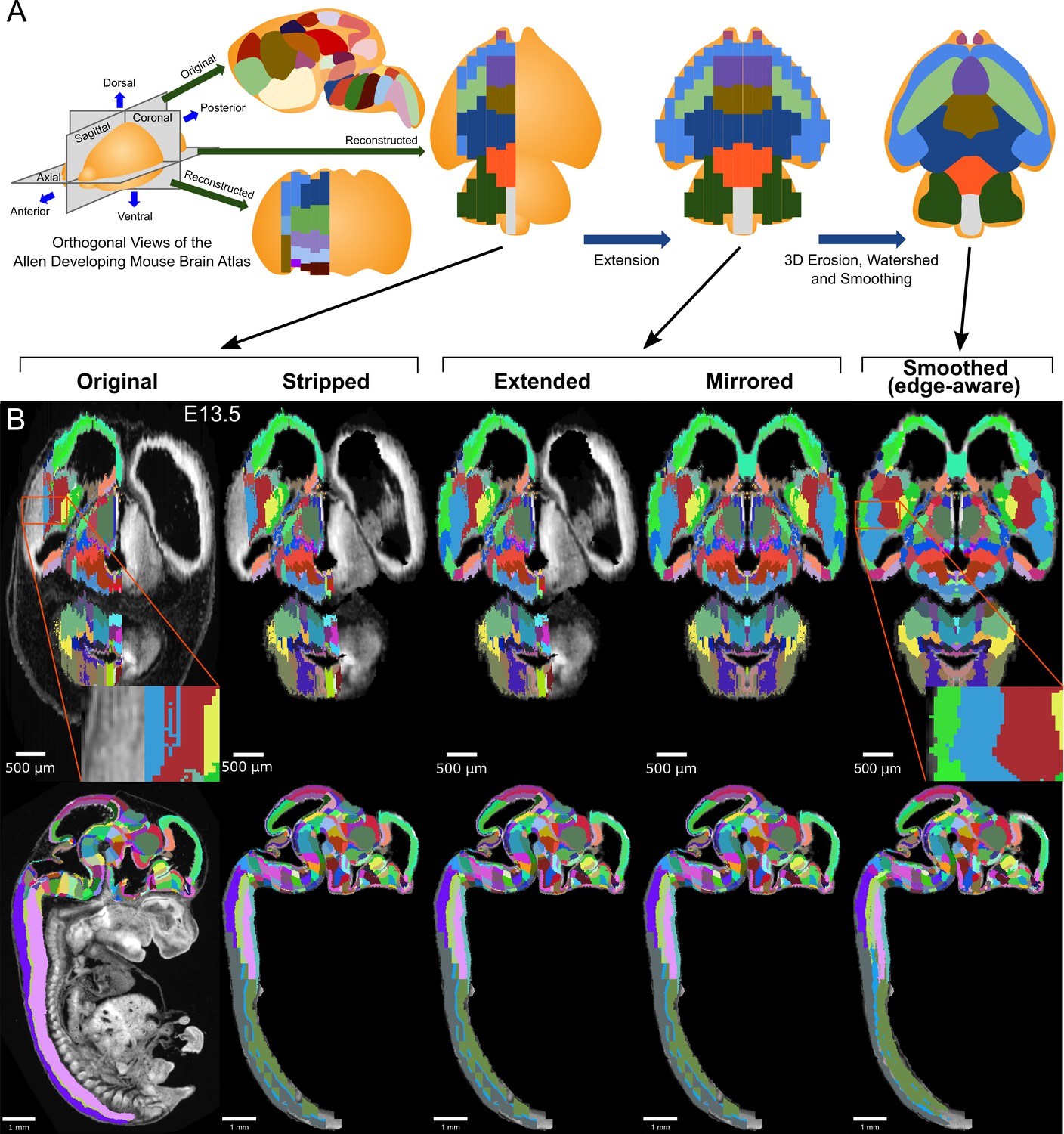
Constructing and optimizing 3D atlases from 2D data with application to the developing mouse brain | eLife

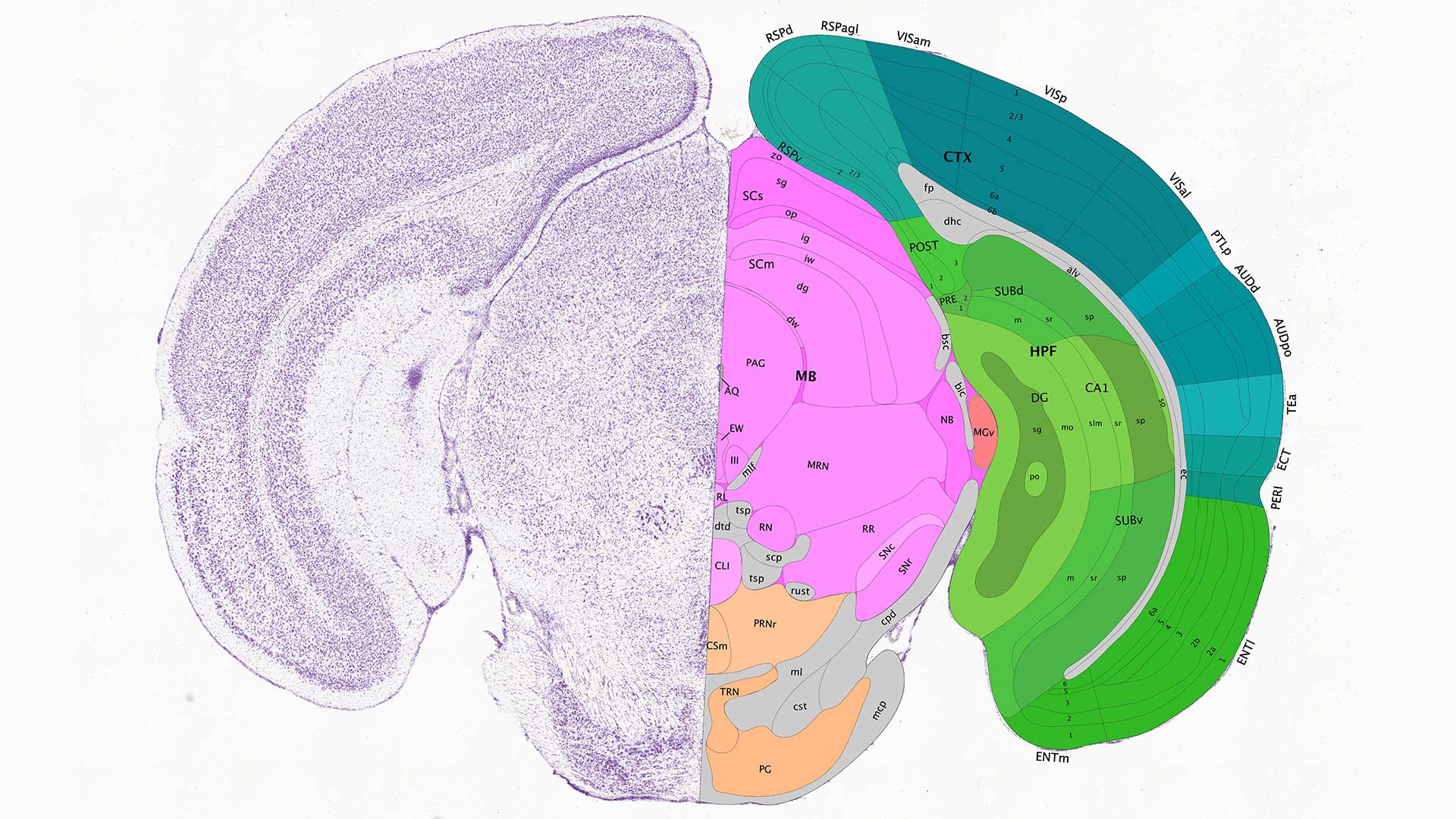
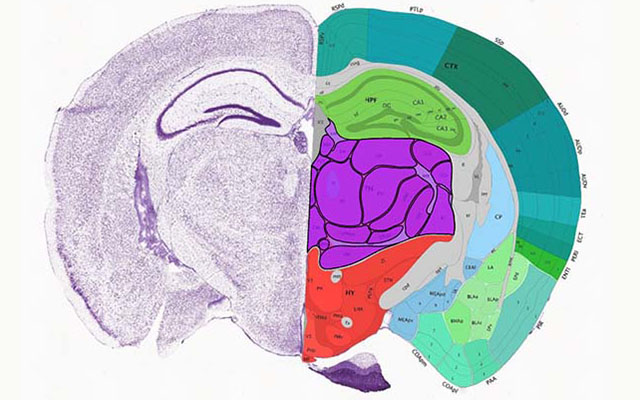
Flexible annotation atlas of the mouse brain: combining and dividing brain structures of the Allen Brain Atlas while maintaining anatomical hierarchy | Scientific Reports